Detail Information of Epigenetic Regulations
| General Information of Drug Transporter (DT) | |||||
|---|---|---|---|---|---|
| DT ID | DTD0114 Transporter Info | ||||
| Gene Name | SLC17A1 | ||||
| Transporter Name | Sodium-dependent phosphate transport protein 1 | ||||
| Gene ID | |||||
| UniProt ID | |||||
| Epigenetic Regulations of This DT (EGR) | |||||
|---|---|---|---|---|---|
|
Methylation |
|||||
|
Atypical teratoid rhabdoid tumor |
4 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in atypical teratoid rhabdoid tumor | [ 1 ] | |||
|
Location |
5'UTR (cg03835296) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.14E+00 | Statistic Test | p-value: 2.03E-08; Z-score: 1.69E+00 | ||
|
Methylation in Case |
9.30E-01 (Median) | Methylation in Control | 8.13E-01 (Median) | ||
|
Studied Phenotype |
Atypical teratoid rhabdoid tumor [ ICD-11: 2A00.1Y] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in atypical teratoid rhabdoid tumor | [ 1 ] | |||
|
Location |
5'UTR (cg22101098) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.04E+00 | Statistic Test | p-value: 5.69E-06; Z-score: -6.23E-01 | ||
|
Methylation in Case |
8.72E-01 (Median) | Methylation in Control | 9.09E-01 (Median) | ||
|
Studied Phenotype |
Atypical teratoid rhabdoid tumor [ ICD-11: 2A00.1Y] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in atypical teratoid rhabdoid tumor | [ 1 ] | |||
|
Location |
5'UTR (cg24766398) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.35E+00 | Statistic Test | p-value: 8.18E-06; Z-score: -1.13E+00 | ||
|
Methylation in Case |
3.24E-01 (Median) | Methylation in Control | 4.38E-01 (Median) | ||
|
Studied Phenotype |
Atypical teratoid rhabdoid tumor [ ICD-11: 2A00.1Y] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 4 |
Methylation of SLC17A1 in atypical teratoid rhabdoid tumor | [ 1 ] | |||
|
Location |
3'UTR (cg06885175) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.10E+00 | Statistic Test | p-value: 1.98E-13; Z-score: -2.85E+00 | ||
|
Methylation in Case |
8.19E-01 (Median) | Methylation in Control | 9.03E-01 (Median) | ||
|
Studied Phenotype |
Atypical teratoid rhabdoid tumor [ ICD-11: 2A00.1Y] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Bladder cancer |
5 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in bladder cancer | [ 2 ] | |||
|
Location |
5'UTR (cg24766398) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -2.40E+00 | Statistic Test | p-value: 7.32E-11; Z-score: -1.04E+01 | ||
|
Methylation in Case |
2.41E-01 (Median) | Methylation in Control | 5.77E-01 (Median) | ||
|
Studied Phenotype |
Bladder cancer [ ICD-11: 2C94] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in bladder cancer | [ 2 ] | |||
|
Location |
5'UTR (cg03835296) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.73E+00 | Statistic Test | p-value: 1.24E-10; Z-score: -2.91E+01 | ||
|
Methylation in Case |
4.76E-01 (Median) | Methylation in Control | 8.23E-01 (Median) | ||
|
Studied Phenotype |
Bladder cancer [ ICD-11: 2C94] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in bladder cancer | [ 2 ] | |||
|
Location |
5'UTR (cg22101098) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.38E+00 | Statistic Test | p-value: 1.25E-07; Z-score: -1.48E+01 | ||
|
Methylation in Case |
5.78E-01 (Median) | Methylation in Control | 7.97E-01 (Median) | ||
|
Studied Phenotype |
Bladder cancer [ ICD-11: 2C94] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 4 |
Methylation of SLC17A1 in bladder cancer | [ 2 ] | |||
|
Location |
TSS200 (cg22988581) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -2.39E+00 | Statistic Test | p-value: 8.47E-06; Z-score: -5.22E+00 | ||
|
Methylation in Case |
1.53E-01 (Median) | Methylation in Control | 3.64E-01 (Median) | ||
|
Studied Phenotype |
Bladder cancer [ ICD-11: 2C94] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 5 |
Methylation of SLC17A1 in bladder cancer | [ 2 ] | |||
|
Location |
3'UTR (cg06885175) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -4.03E+00 | Statistic Test | p-value: 3.19E-05; Z-score: -5.52E+00 | ||
|
Methylation in Case |
8.50E-02 (Median) | Methylation in Control | 3.42E-01 (Median) | ||
|
Studied Phenotype |
Bladder cancer [ ICD-11: 2C94] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Breast cancer |
4 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in breast cancer | [ 3 ] | |||
|
Location |
5'UTR (cg22101098) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.15E+00 | Statistic Test | p-value: 1.06E-05; Z-score: 1.77E+00 | ||
|
Methylation in Case |
7.74E-01 (Median) | Methylation in Control | 6.71E-01 (Median) | ||
|
Studied Phenotype |
Breast cancer [ ICD-11: 2C60-2C6Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in breast cancer | [ 3 ] | |||
|
Location |
5'UTR (cg24766398) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.15E+00 | Statistic Test | p-value: 2.37E-02; Z-score: 8.86E-01 | ||
|
Methylation in Case |
4.63E-01 (Median) | Methylation in Control | 4.03E-01 (Median) | ||
|
Studied Phenotype |
Breast cancer [ ICD-11: 2C60-2C6Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in breast cancer | [ 3 ] | |||
|
Location |
TSS200 (cg22988581) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.09E+00 | Statistic Test | p-value: 2.17E-03; Z-score: -9.72E-01 | ||
|
Methylation in Case |
4.65E-01 (Median) | Methylation in Control | 5.08E-01 (Median) | ||
|
Studied Phenotype |
Breast cancer [ ICD-11: 2C60-2C6Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 4 |
Methylation of SLC17A1 in breast cancer | [ 3 ] | |||
|
Location |
Body (cg19490609) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.29E+00 | Statistic Test | p-value: 7.45E-07; Z-score: 1.83E+00 | ||
|
Methylation in Case |
4.59E-01 (Median) | Methylation in Control | 3.56E-01 (Median) | ||
|
Studied Phenotype |
Breast cancer [ ICD-11: 2C60-2C6Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Colorectal cancer |
6 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in colorectal cancer | [ 4 ] | |||
|
Location |
5'UTR (cg24766398) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.23E+00 | Statistic Test | p-value: 1.93E-07; Z-score: -1.82E+00 | ||
|
Methylation in Case |
6.30E-01 (Median) | Methylation in Control | 7.74E-01 (Median) | ||
|
Studied Phenotype |
Colorectal cancer [ ICD-11: 2B91] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in colorectal cancer | [ 4 ] | |||
|
Location |
5'UTR (cg03835296) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.05E+00 | Statistic Test | p-value: 1.21E-06; Z-score: -2.15E+00 | ||
|
Methylation in Case |
8.68E-01 (Median) | Methylation in Control | 9.11E-01 (Median) | ||
|
Studied Phenotype |
Colorectal cancer [ ICD-11: 2B91] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in colorectal cancer | [ 4 ] | |||
|
Location |
5'UTR (cg22101098) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.03E+00 | Statistic Test | p-value: 1.22E-04; Z-score: -9.24E-01 | ||
|
Methylation in Case |
8.78E-01 (Median) | Methylation in Control | 9.06E-01 (Median) | ||
|
Studied Phenotype |
Colorectal cancer [ ICD-11: 2B91] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 4 |
Methylation of SLC17A1 in colorectal cancer | [ 4 ] | |||
|
Location |
TSS1500 (cg26691898) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.04E+00 | Statistic Test | p-value: 4.41E-07; Z-score: -2.07E+00 | ||
|
Methylation in Case |
8.86E-01 (Median) | Methylation in Control | 9.17E-01 (Median) | ||
|
Studied Phenotype |
Colorectal cancer [ ICD-11: 2B91] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 5 |
Methylation of SLC17A1 in colorectal cancer | [ 4 ] | |||
|
Location |
3'UTR (cg06885175) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.35E+00 | Statistic Test | p-value: 2.38E-04; Z-score: -1.06E+00 | ||
|
Methylation in Case |
2.42E-01 (Median) | Methylation in Control | 3.27E-01 (Median) | ||
|
Studied Phenotype |
Colorectal cancer [ ICD-11: 2B91] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 6 |
Significant hypermethylation of SLC17A1 in colorectal cancer than that in healthy individual | ||||
Studied Phenotype |
Colorectal cancer [ICD-11:2B91] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 6.39E-08; Fold-change: 0.317341629; Z-score: 1.209543568 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue adjacent to the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
HIV infection |
3 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in HIV infection | [ 5 ] | |||
|
Location |
5'UTR (cg24766398) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.07E+00 | Statistic Test | p-value: 2.40E-03; Z-score: -9.90E-01 | ||
|
Methylation in Case |
7.74E-01 (Median) | Methylation in Control | 8.27E-01 (Median) | ||
|
Studied Phenotype |
HIV infection [ ICD-11: 1C62.Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in HIV infection | [ 5 ] | |||
|
Location |
TSS200 (cg22988581) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.04E+00 | Statistic Test | p-value: 1.25E-03; Z-score: -7.84E-01 | ||
|
Methylation in Case |
7.51E-01 (Median) | Methylation in Control | 7.79E-01 (Median) | ||
|
Studied Phenotype |
HIV infection [ ICD-11: 1C62.Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in HIV infection | [ 5 ] | |||
|
Location |
3'UTR (cg06885175) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.51E+00 | Statistic Test | p-value: 1.24E-04; Z-score: 1.35E+00 | ||
|
Methylation in Case |
2.52E-01 (Median) | Methylation in Control | 1.67E-01 (Median) | ||
|
Studied Phenotype |
HIV infection [ ICD-11: 1C62.Z] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Lung adenocarcinoma |
2 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in lung adenocarcinoma | [ 6 ] | |||
|
Location |
5'UTR (cg03835296) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.08E+00 | Statistic Test | p-value: 1.56E-02; Z-score: -1.30E+00 | ||
|
Methylation in Case |
7.90E-01 (Median) | Methylation in Control | 8.52E-01 (Median) | ||
|
Studied Phenotype |
Lung adenocarcinoma [ ICD-11: 2C25.0] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in lung adenocarcinoma | [ 6 ] | |||
|
Location |
TSS200 (cg22988581) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.20E+00 | Statistic Test | p-value: 2.67E-02; Z-score: -1.63E+00 | ||
|
Methylation in Case |
5.31E-01 (Median) | Methylation in Control | 6.35E-01 (Median) | ||
|
Studied Phenotype |
Lung adenocarcinoma [ ICD-11: 2C25.0] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Pancretic ductal adenocarcinoma |
4 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in pancretic ductal adenocarcinoma | [ 7 ] | |||
|
Location |
5'UTR (cg05043886) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.35E+00 | Statistic Test | p-value: 2.23E-09; Z-score: 1.71E+00 | ||
|
Methylation in Case |
3.25E-01 (Median) | Methylation in Control | 2.41E-01 (Median) | ||
|
Studied Phenotype |
Pancretic ductal adenocarcinoma [ ICD-11: 2C10.0] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in pancretic ductal adenocarcinoma | [ 7 ] | |||
|
Location |
TSS1500 (cg01151699) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.36E+00 | Statistic Test | p-value: 2.43E-14; Z-score: 1.59E+00 | ||
|
Methylation in Case |
7.54E-02 (Median) | Methylation in Control | 5.52E-02 (Median) | ||
|
Studied Phenotype |
Pancretic ductal adenocarcinoma [ ICD-11: 2C10.0] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in pancretic ductal adenocarcinoma | [ 7 ] | |||
|
Location |
TSS1500 (cg00999623) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.19E+00 | Statistic Test | p-value: 2.30E-04; Z-score: -8.97E-01 | ||
|
Methylation in Case |
1.32E-01 (Median) | Methylation in Control | 1.58E-01 (Median) | ||
|
Studied Phenotype |
Pancretic ductal adenocarcinoma [ ICD-11: 2C10.0] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 4 |
Methylation of SLC17A1 in pancretic ductal adenocarcinoma | [ 7 ] | |||
|
Location |
TSS1500 (cg08424219) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.24E+00 | Statistic Test | p-value: 8.34E-03; Z-score: -4.52E-01 | ||
|
Methylation in Case |
9.65E-02 (Median) | Methylation in Control | 1.19E-01 (Median) | ||
|
Studied Phenotype |
Pancretic ductal adenocarcinoma [ ICD-11: 2C10.0] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Papillary thyroid cancer |
2 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in papillary thyroid cancer | [ 8 ] | |||
|
Location |
5'UTR (cg22101098) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.11E+00 | Statistic Test | p-value: 2.47E-03; Z-score: 6.92E-01 | ||
|
Methylation in Case |
5.68E-01 (Median) | Methylation in Control | 5.11E-01 (Median) | ||
|
Studied Phenotype |
Papillary thyroid cancer [ ICD-11: 2D10.1] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in papillary thyroid cancer | [ 8 ] | |||
|
Location |
3'UTR (cg06885175) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.31E+00 | Statistic Test | p-value: 5.36E-04; Z-score: -1.19E+00 | ||
|
Methylation in Case |
2.48E-01 (Median) | Methylation in Control | 3.26E-01 (Median) | ||
|
Studied Phenotype |
Papillary thyroid cancer [ ICD-11: 2D10.1] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Hepatocellular carcinoma |
5 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Methylation of SLC17A1 in hepatocellular carcinoma | [ 9 ] | |||
|
Location |
TSS1500 (cg26691898) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.03E+00 | Statistic Test | p-value: 2.37E-04; Z-score: -6.36E-01 | ||
|
Methylation in Case |
7.88E-01 (Median) | Methylation in Control | 8.09E-01 (Median) | ||
|
Studied Phenotype |
Hepatocellular carcinoma [ ICD-11: 2C12.02] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 2 |
Methylation of SLC17A1 in hepatocellular carcinoma | [ 9 ] | |||
|
Location |
TSS200 (cg07888856) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 6.70E+00 | Statistic Test | p-value: 9.38E-14; Z-score: 1.31E+01 | ||
|
Methylation in Case |
2.76E-01 (Median) | Methylation in Control | 4.12E-02 (Median) | ||
|
Studied Phenotype |
Hepatocellular carcinoma [ ICD-11: 2C12.02] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 3 |
Methylation of SLC17A1 in hepatocellular carcinoma | [ 9 ] | |||
|
Location |
TSS200 (cg07808555) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 2.46E+00 | Statistic Test | p-value: 5.08E-12; Z-score: 2.49E+00 | ||
|
Methylation in Case |
1.85E-01 (Median) | Methylation in Control | 7.54E-02 (Median) | ||
|
Studied Phenotype |
Hepatocellular carcinoma [ ICD-11: 2C12.02] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 4 |
Methylation of SLC17A1 in hepatocellular carcinoma | [ 9 ] | |||
|
Location |
Body (cg19490609) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: 1.11E+00 | Statistic Test | p-value: 3.02E-02; Z-score: 8.78E-01 | ||
|
Methylation in Case |
6.86E-01 (Median) | Methylation in Control | 6.17E-01 (Median) | ||
|
Studied Phenotype |
Hepatocellular carcinoma [ ICD-11: 2C12.02] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Epigenetic Phenomenon 5 |
Methylation of SLC17A1 in hepatocellular carcinoma | [ 9 ] | |||
|
Location |
3'UTR (cg06885175) | ||||
|
Epigenetic Type |
Methylation | Experiment Method | Infinium HumanMethylation450 BeadChip | ||
|
Methylation Fold Change |
Fold Change: -1.25E+00 | Statistic Test | p-value: 6.70E-05; Z-score: -7.98E-01 | ||
|
Methylation in Case |
1.32E-01 (Median) | Methylation in Control | 1.66E-01 (Median) | ||
|
Studied Phenotype |
Hepatocellular carcinoma [ ICD-11: 2C12.02] | ||||
|
Experimental Material |
Patient tissue samples | ||||
|
Liver cancer |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
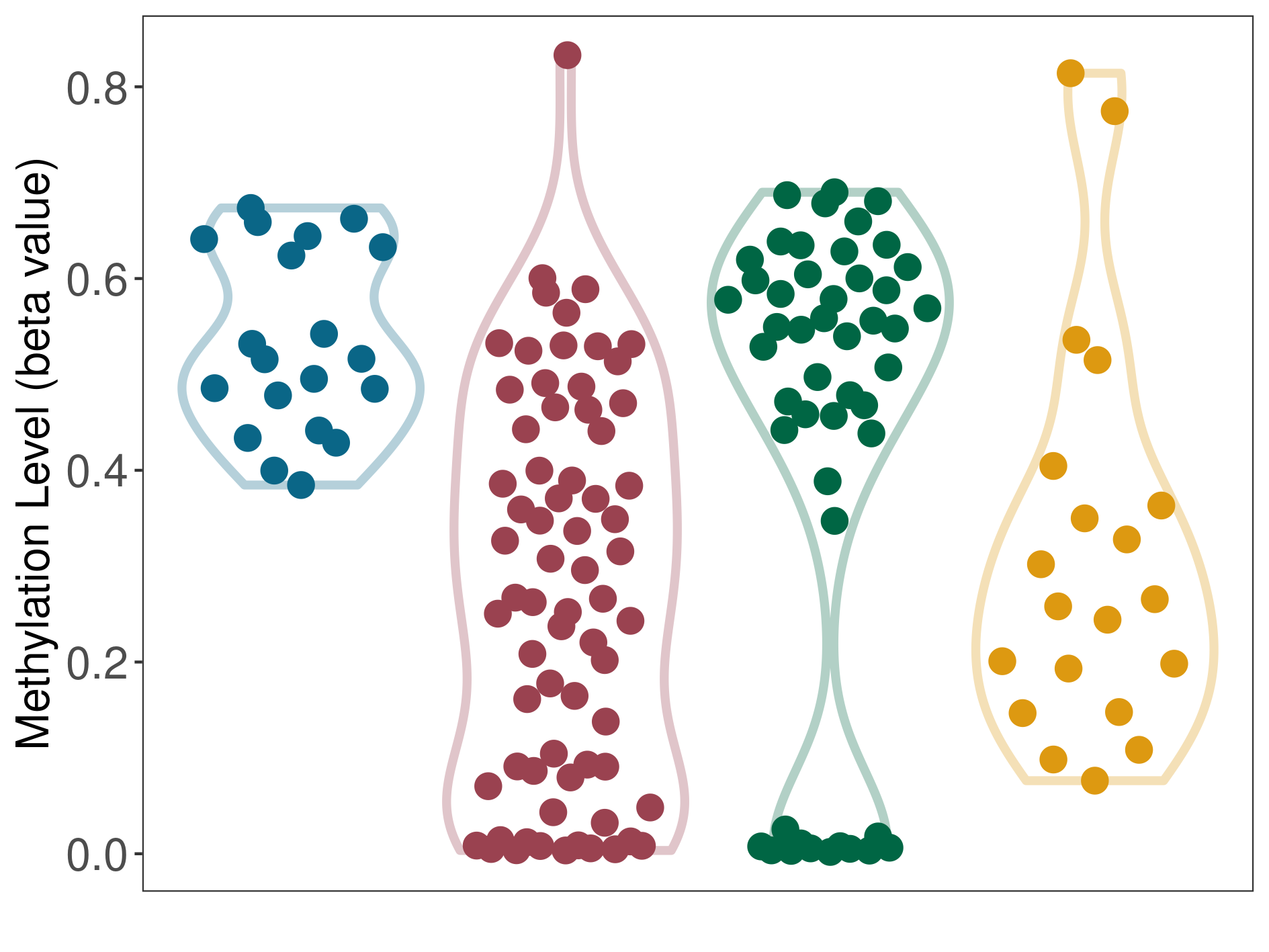
Moderate/moderate hypomethylation of SLC17A1 in liver cancer than that in healthy individual/adjacent tissue | ||||
Studied Phenotype |
Liver cancer [ICD-11:2C12] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.000857727; Fold-change: -0.273796368; Z-score: -1.094861211 | ||||
The Methylation Level of Disease Section Compare with the Adjacent Tissue |
p-value: 2.07E-11; Fold-change: -0.250044978; Z-score: -2.619104924 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue adjacent to the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
DT methylation level in tissue other than the diseased tissue of patients
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Cerebellar liponeurocytoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
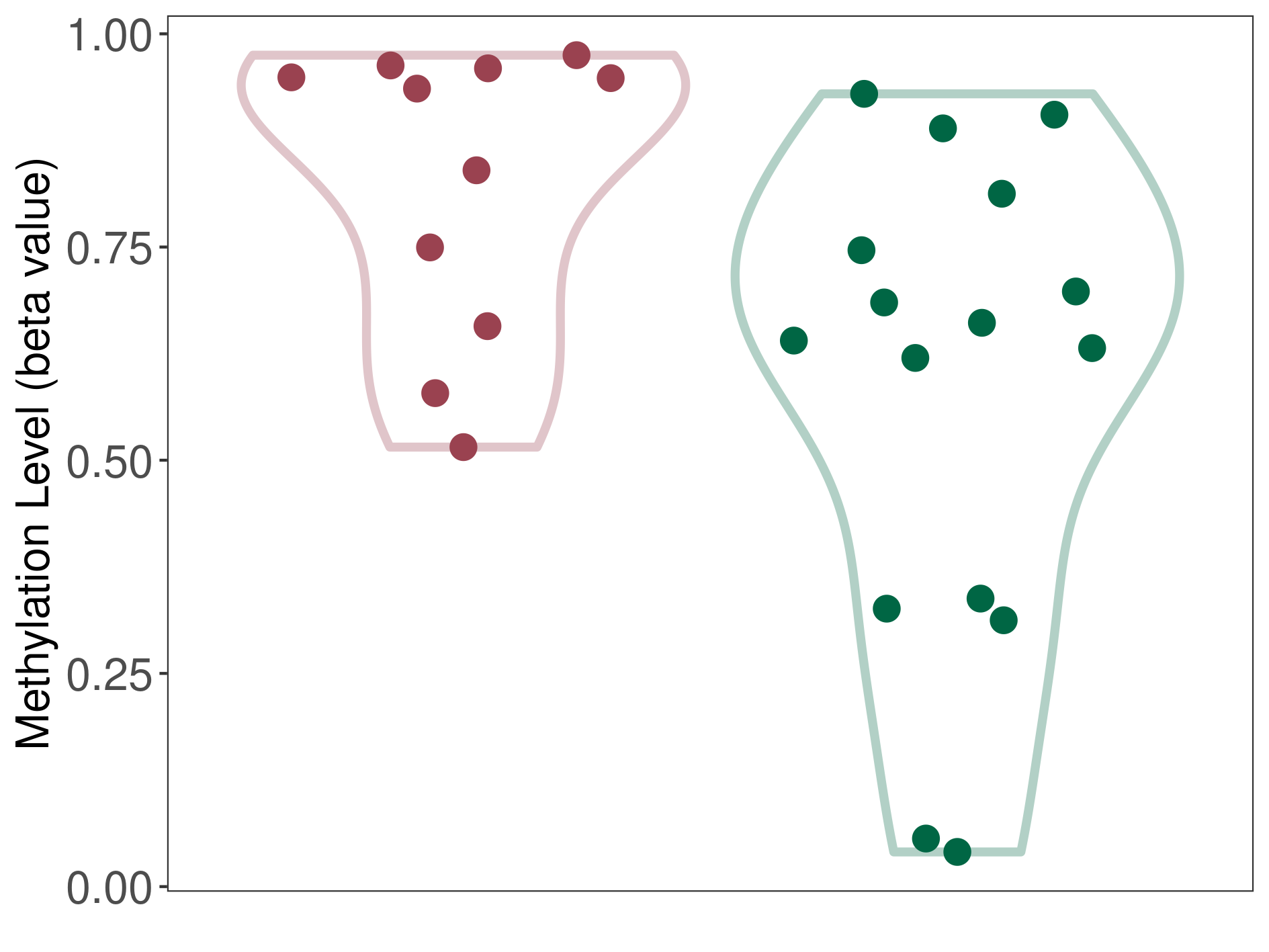
Moderate hypermethylation of SLC17A1 in cerebellar liponeurocytoma than that in healthy individual | ||||
Studied Phenotype |
Cerebellar liponeurocytoma [ICD-11:2A00.0Y] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.010171838; Fold-change: 0.285047126; Z-score: 1.005873949 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Oligodendroglioma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
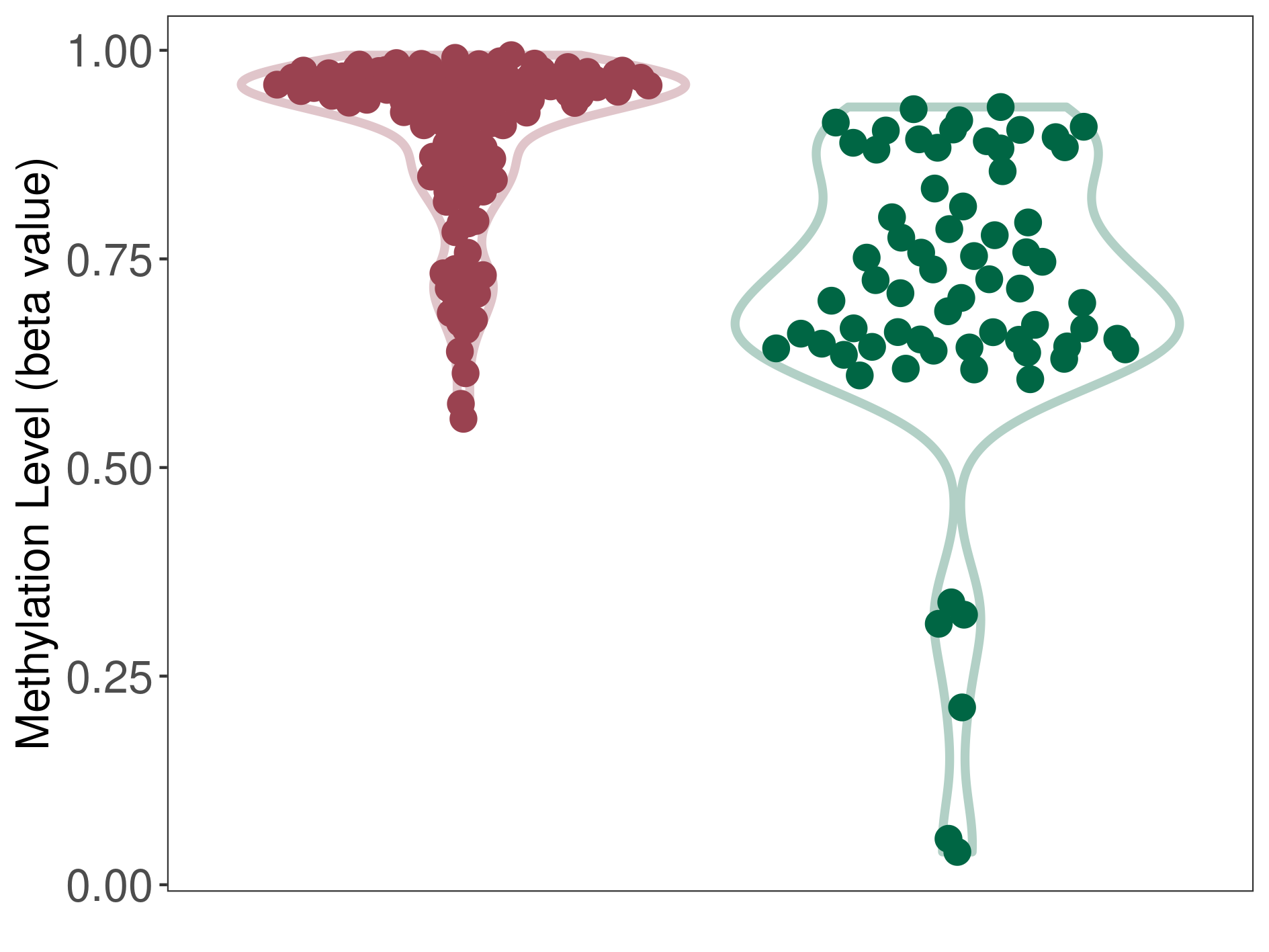
Moderate hypermethylation of SLC17A1 in oligodendroglioma than that in healthy individual | ||||
Studied Phenotype |
Oligodendroglioma [ICD-11:2A00.0Y] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 5.48E-12; Fold-change: 0.23651157; Z-score: 1.25327646 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
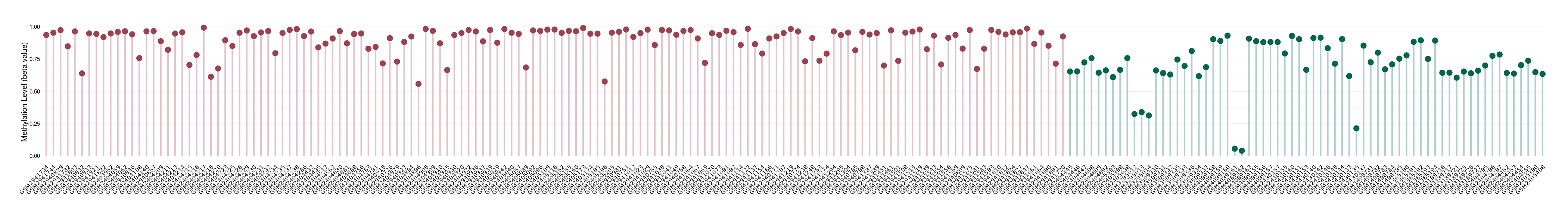 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Pituitary adenoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
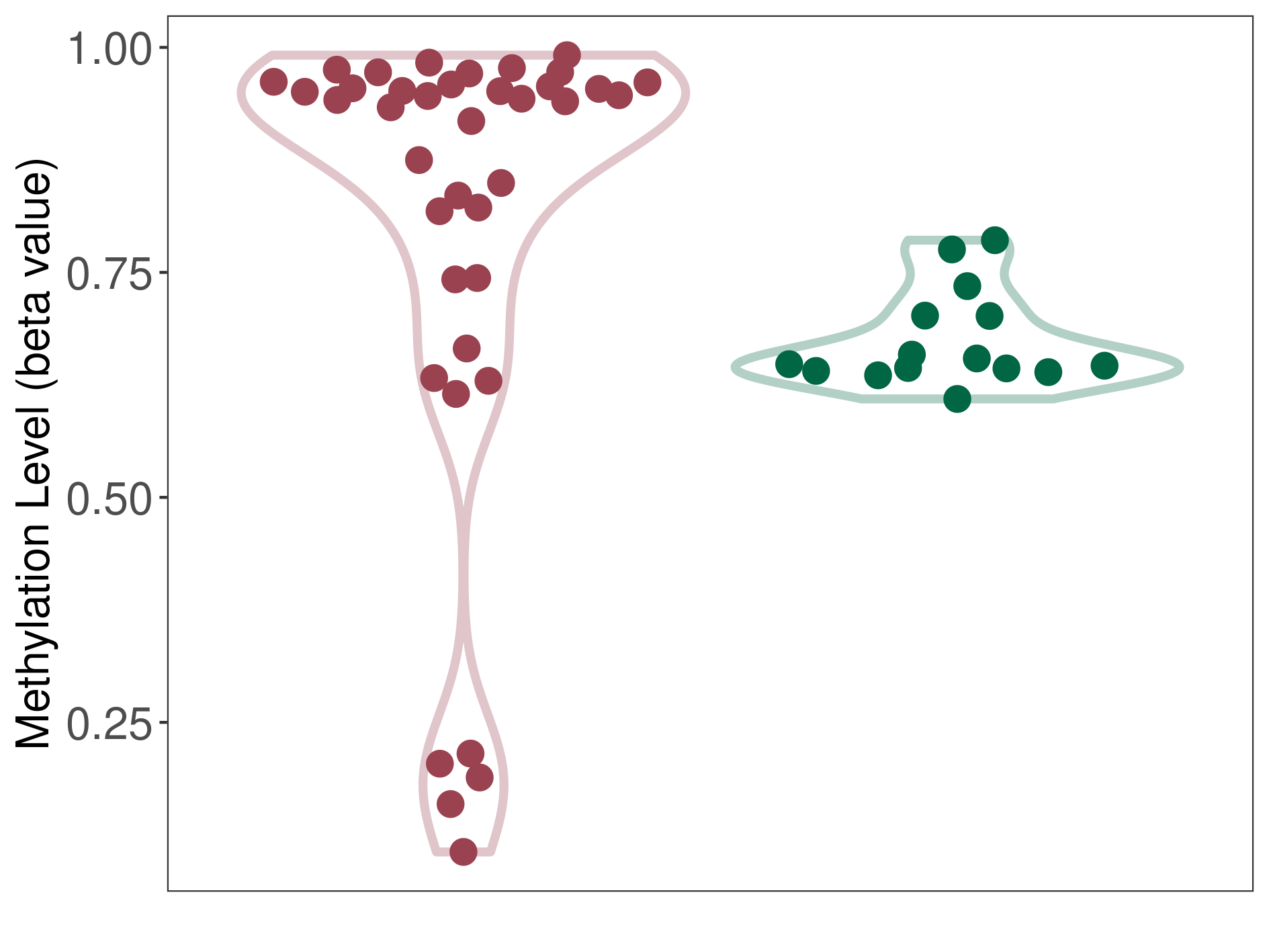
Moderate hypermethylation of SLC17A1 in pituitary adenoma than that in healthy individual | ||||
Studied Phenotype |
Pituitary adenoma [ICD-11:2F37] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.008230396; Fold-change: 0.293560842; Z-score: 5.497815096 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Chordoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
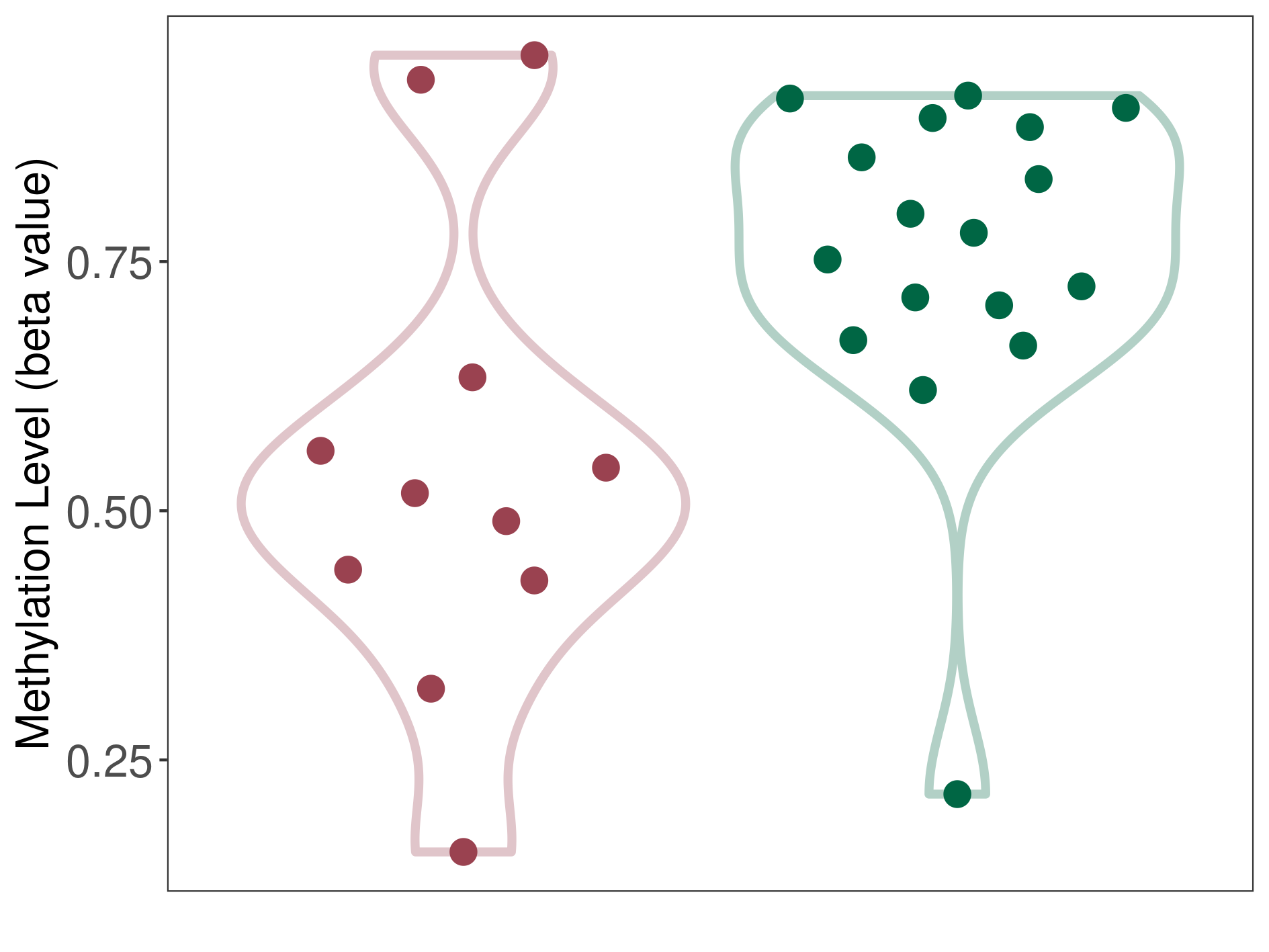
Moderate hypomethylation of SLC17A1 in chordoma than that in healthy individual | ||||
Studied Phenotype |
Chordoma [ICD-11:5A61.0] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.019679349; Fold-change: -0.261206131; Z-score: -1.545974161 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Craniopharyngioma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
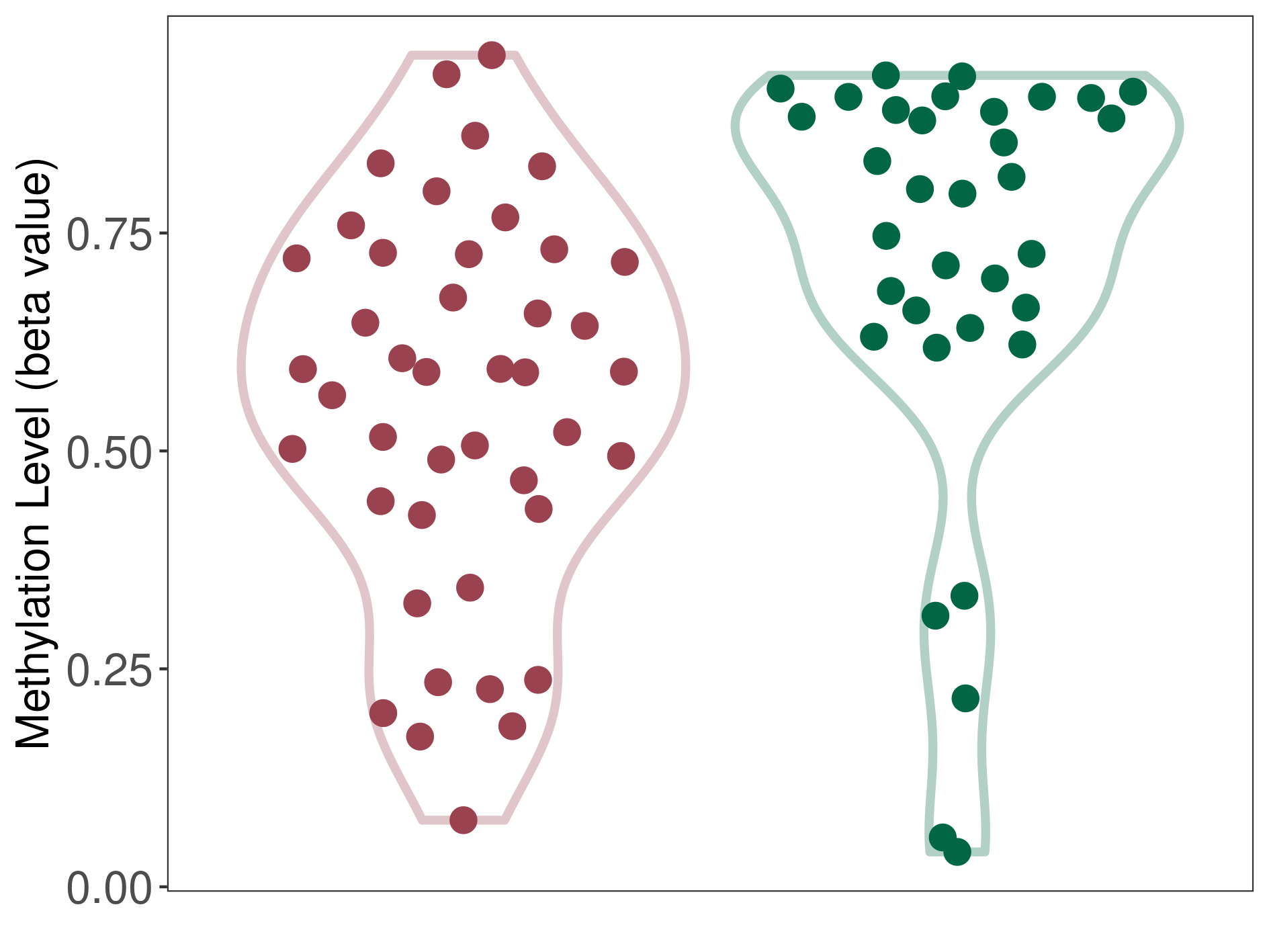
Moderate hypomethylation of SLC17A1 in craniopharyngioma than that in healthy individual | ||||
Studied Phenotype |
Craniopharyngioma [ICD-11:2F9A] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.005118004; Fold-change: -0.207162332; Z-score: -0.840836639 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
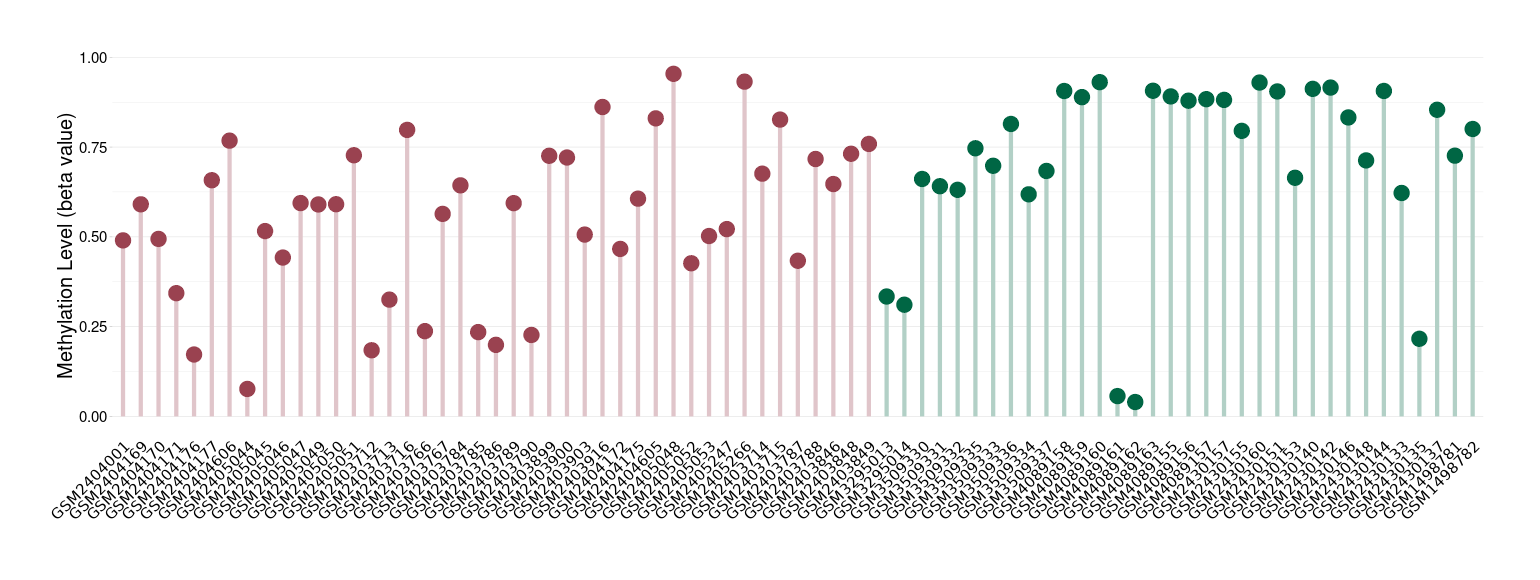 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Diffuse midline glioma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
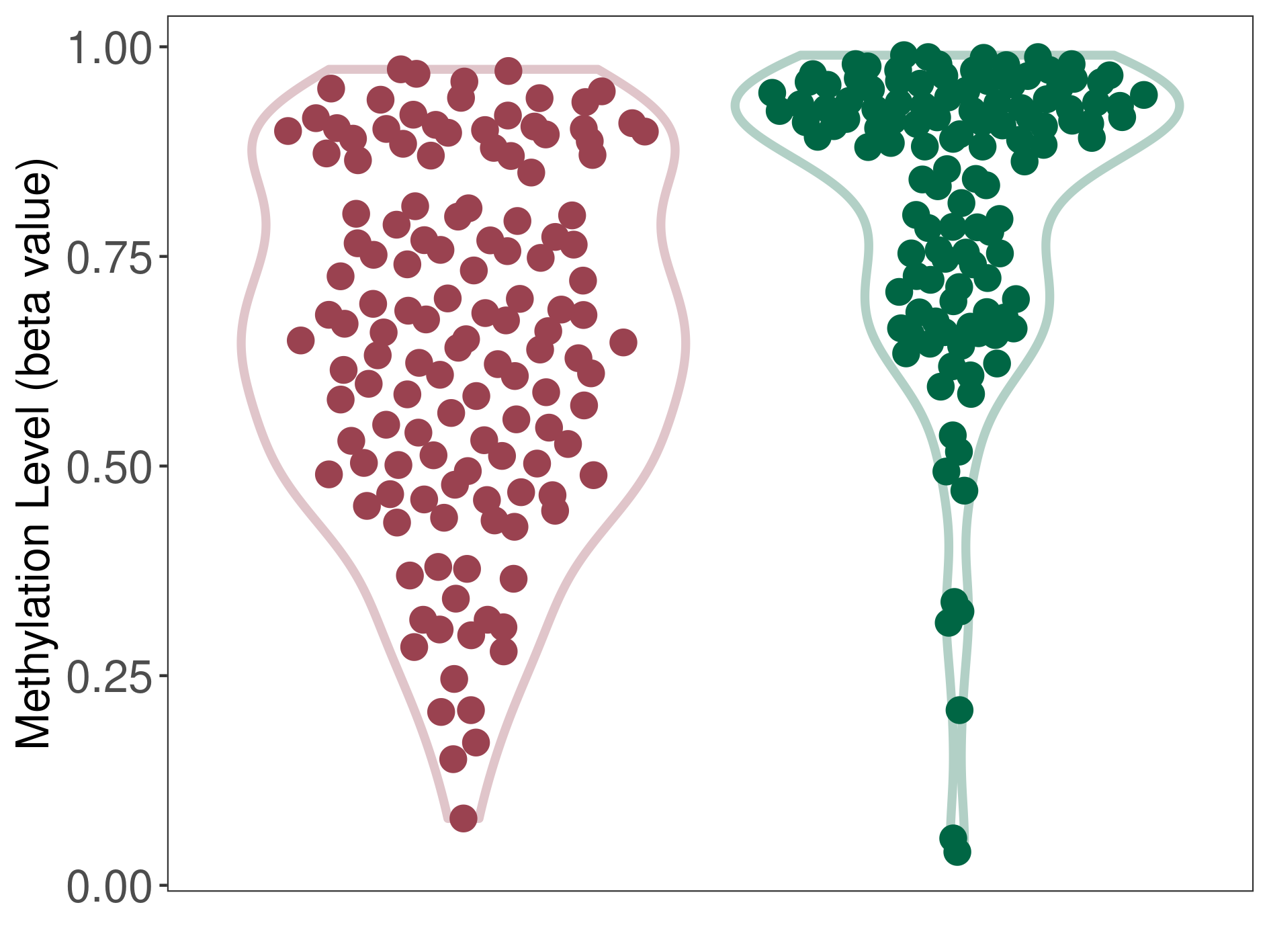
Moderate hypomethylation of SLC17A1 in diffuse midline glioma than that in healthy individual | ||||
Studied Phenotype |
Diffuse midline glioma [ICD-11:2A00.0Z] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 1.02E-09; Fold-change: -0.230624986; Z-score: -1.222796419 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Osteoarthritis |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
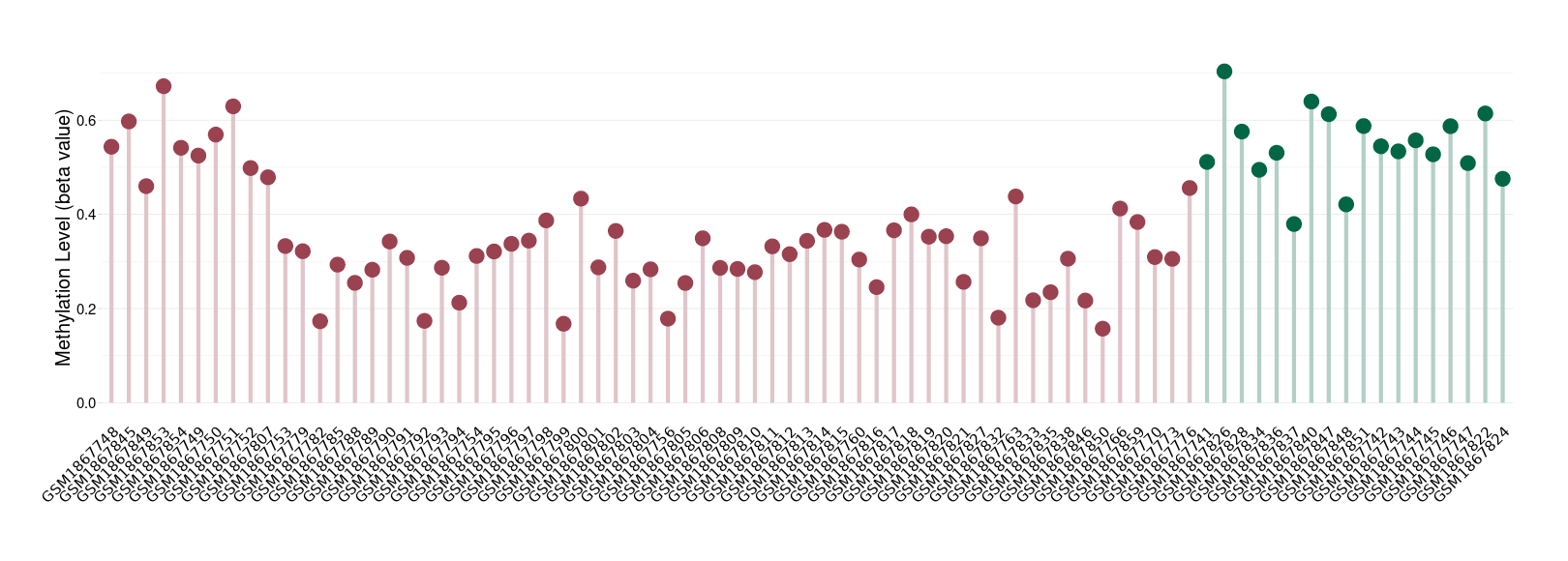
Moderate hypomethylation of SLC17A1 in osteoarthritis than that in healthy individual | ||||
Studied Phenotype |
Osteoarthritis [ICD-11:FA00] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 7.93E-11; Fold-change: -0.217409842; Z-score: -2.816531123 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||

|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Asthma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
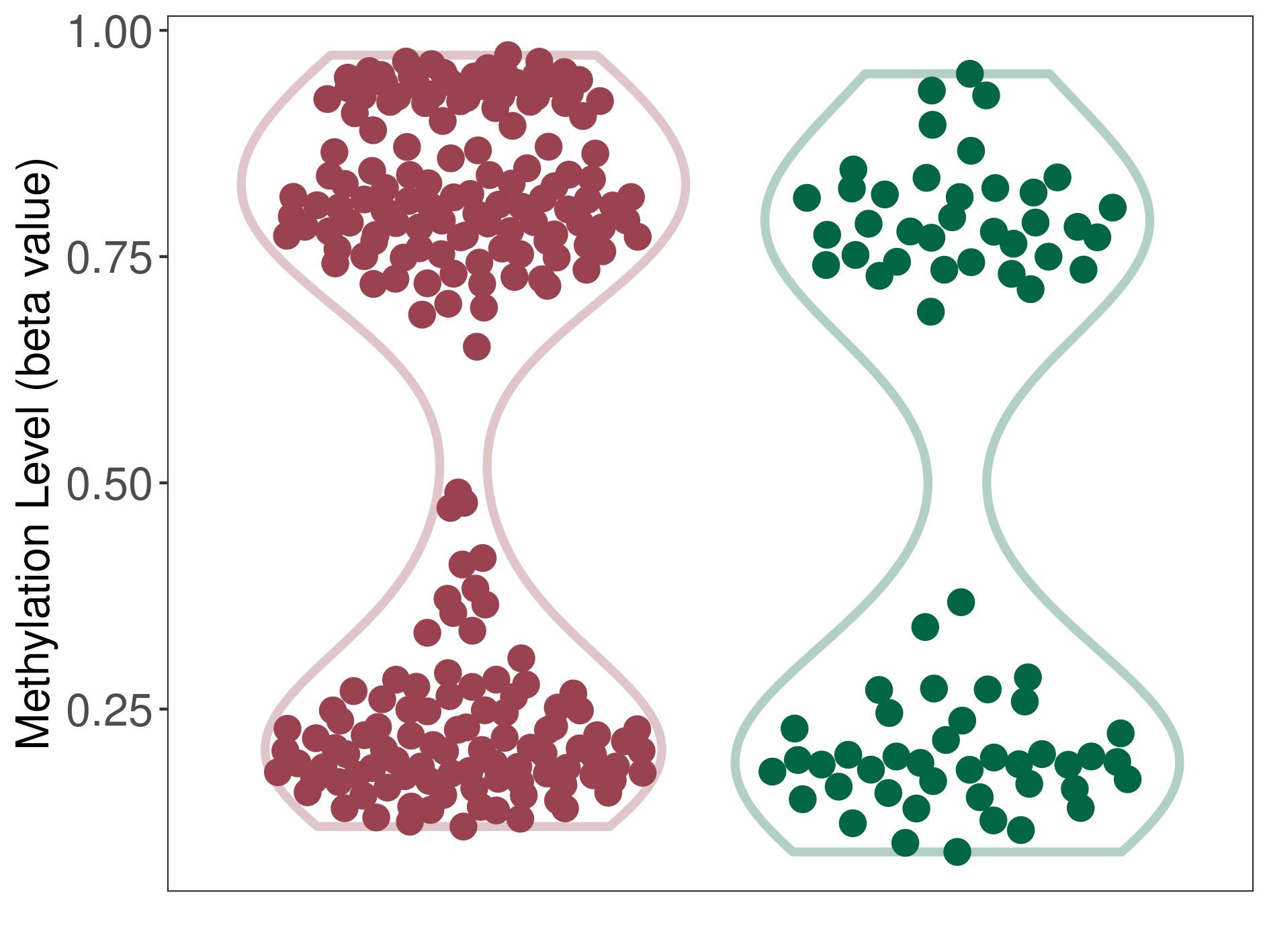
Significant hypermethylation of SLC17A1 in asthma than that in healthy individual | ||||
Studied Phenotype |
Asthma [ICD-11:CA23] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.032594569; Fold-change: 0.45039513; Z-score: 1.464816356 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Prostate cancer metastasis |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
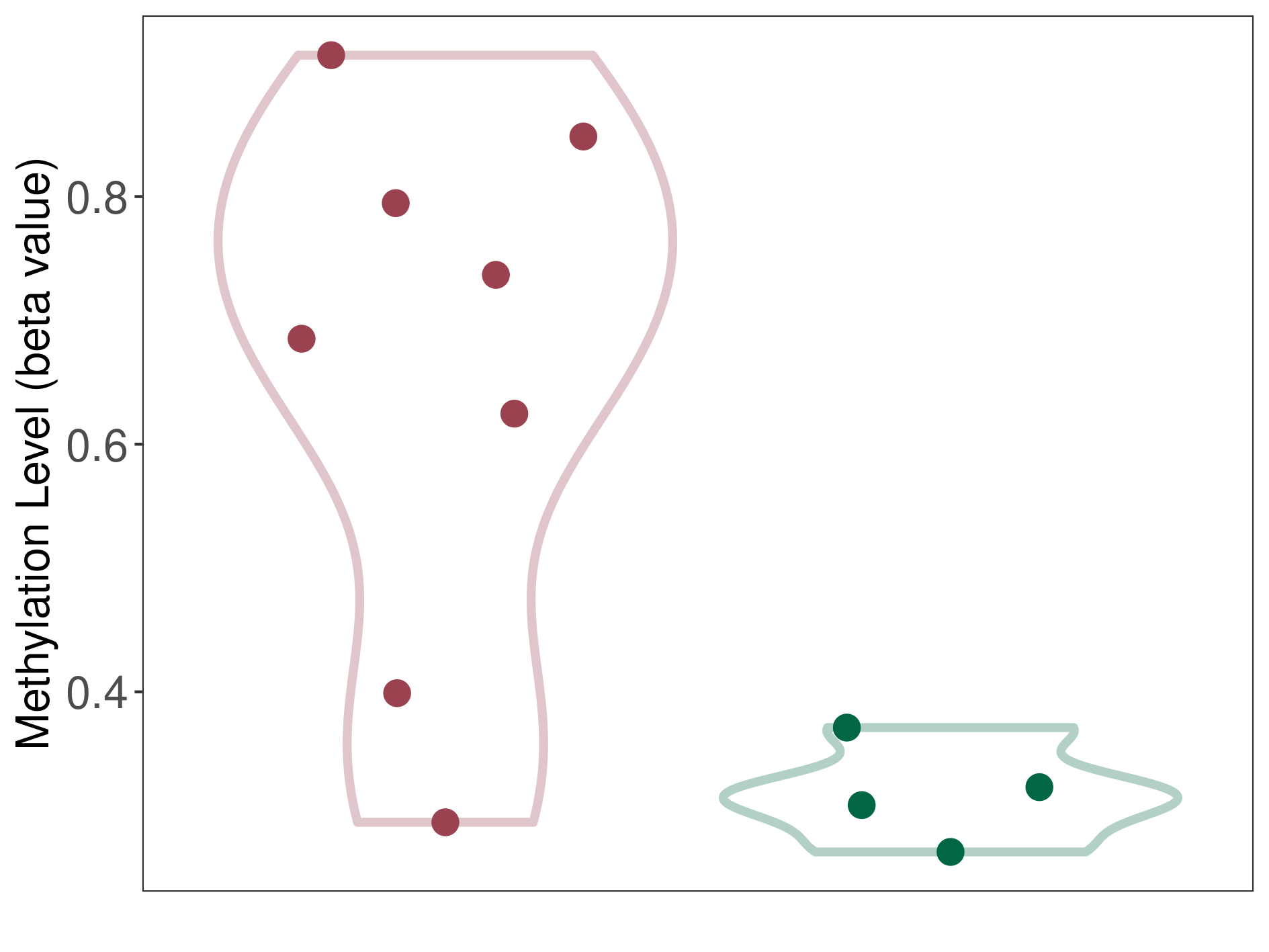
Significant hypermethylation of SLC17A1 in prostate cancer metastasis than that in healthy individual | ||||
Studied Phenotype |
Prostate cancer metastasis [ICD-11:2.00E+06] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.002507466; Fold-change: 0.395182376; Z-score: 9.51586899 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Prostate cancer |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Significant hypermethylation of SLC17A1 in prostate cancer than that in healthy individual | ||||
Studied Phenotype |
Prostate cancer [ICD-11:2C82] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 9.05E-08; Fold-change: 0.663134939; Z-score: 4.549189725 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||

|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Brain neuroblastoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
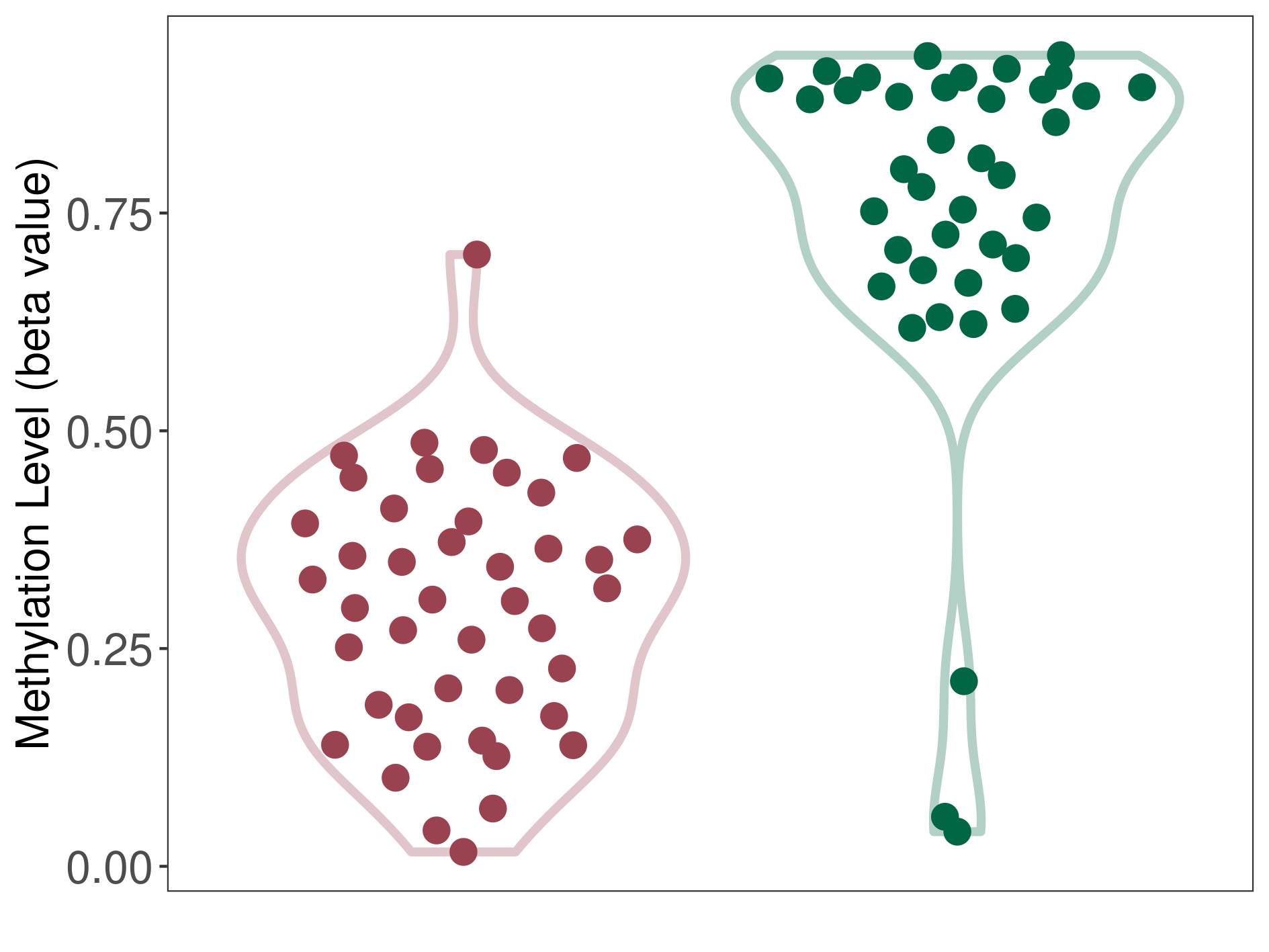
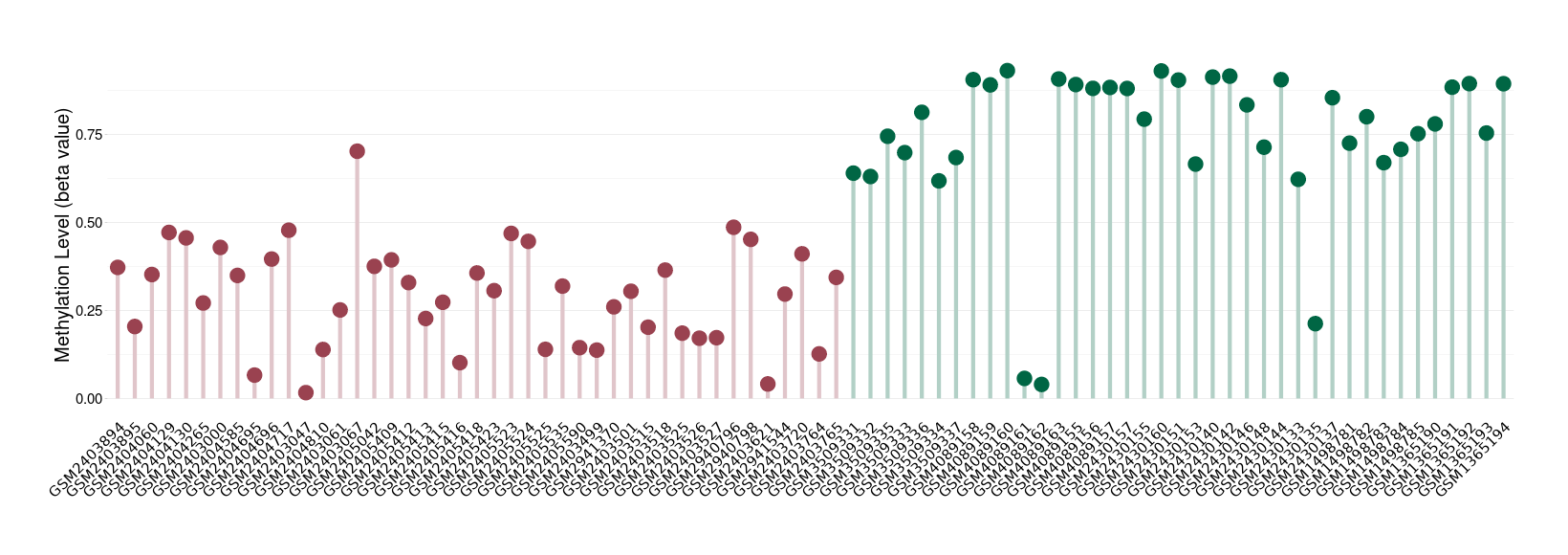
Significant hypomethylation of SLC17A1 in brain neuroblastoma than that in healthy individual | ||||
Studied Phenotype |
Brain neuroblastoma [ICD-11:2A00.11] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 1.22E-16; Fold-change: -0.494165435; Z-score: -2.300695929 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Brain neuroepithelial tumour |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Significant hypomethylation of SLC17A1 in brain neuroepithelial tumour than that in healthy individual | ||||
Studied Phenotype |
Brain neuroepithelial tumour [ICD-11:2A00.2Y] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 4.48E-11; Fold-change: -0.455198117; Z-score: -2.222922786 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||

|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Lymphoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
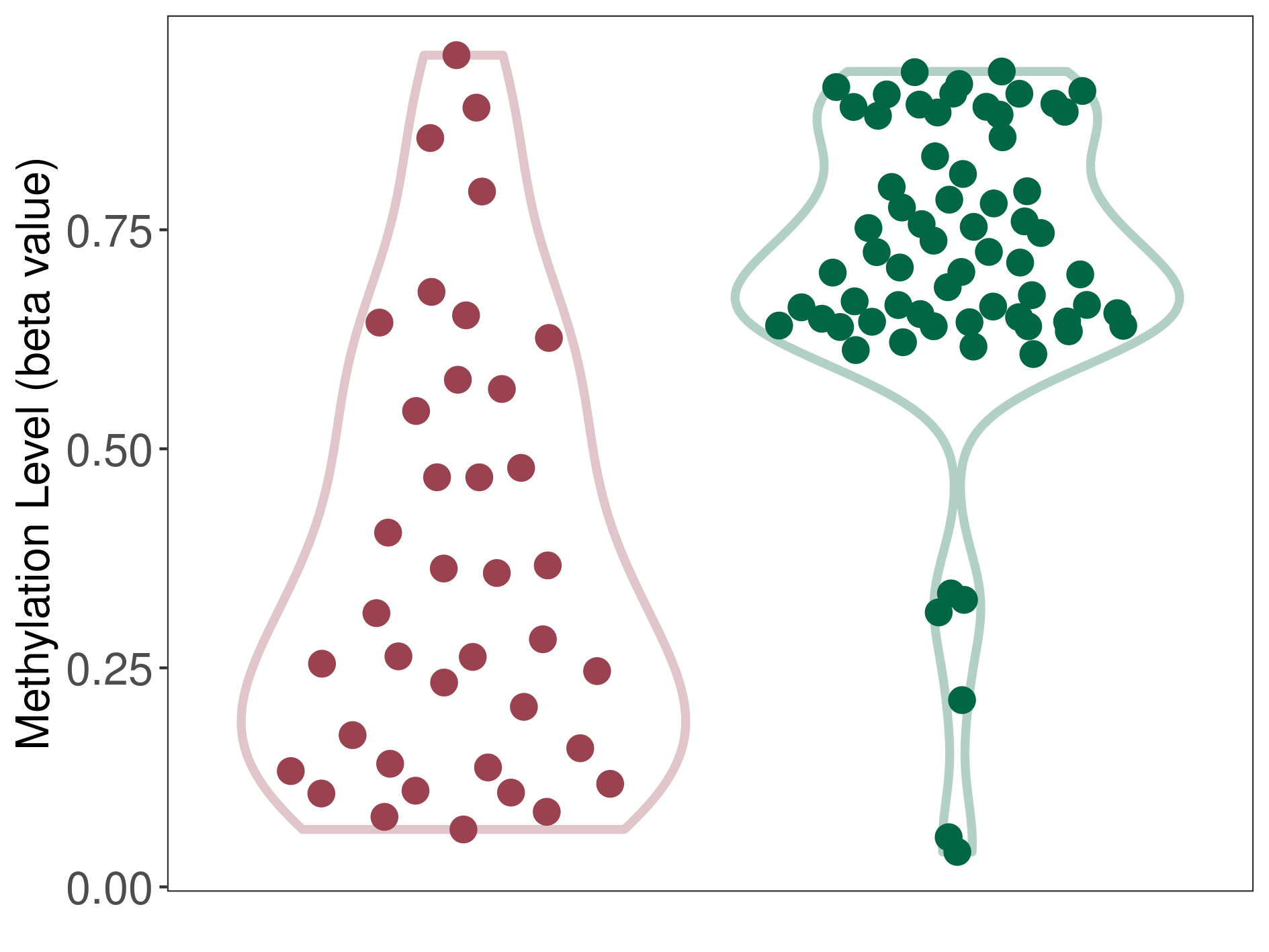
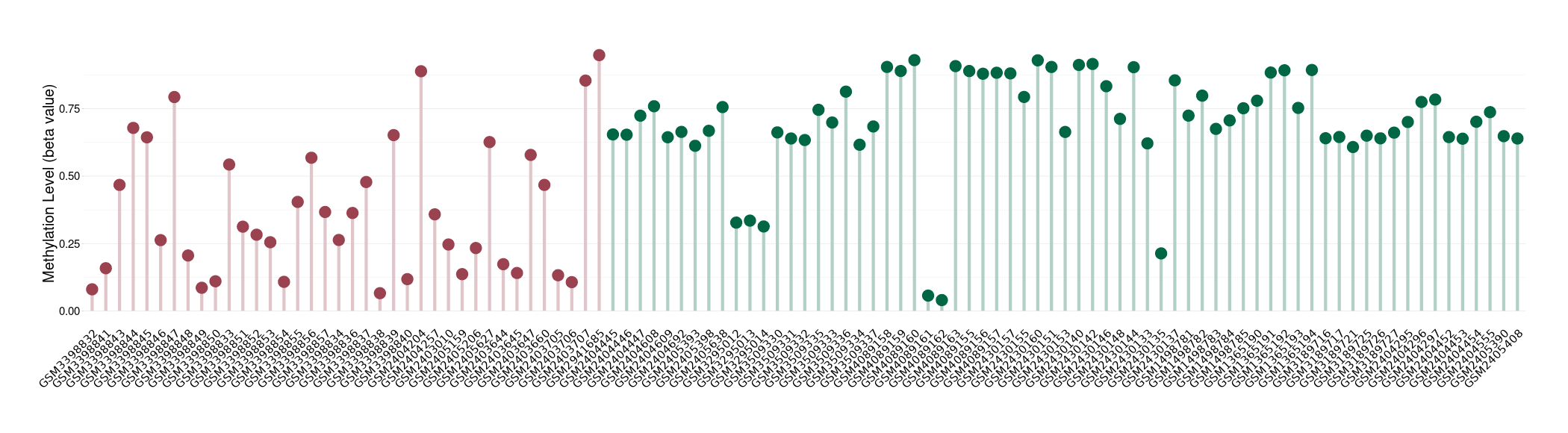
Significant hypomethylation of SLC17A1 in lymphoma than that in healthy individual | ||||
Studied Phenotype |
Lymphoma [ICD-11:2B30] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 2.09E-09; Fold-change: -0.409403034; Z-score: -2.172968245 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Medulloblastoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
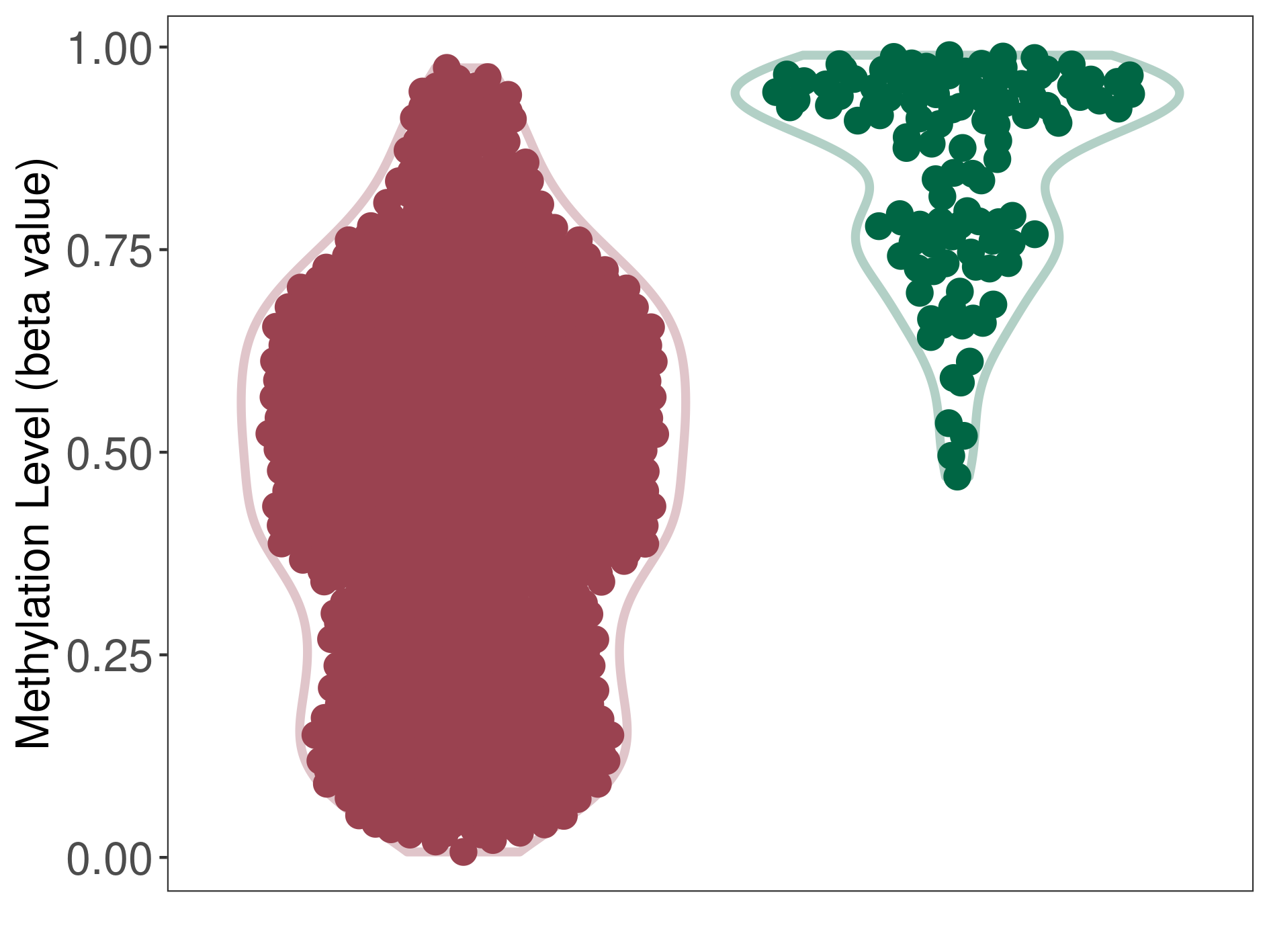
Significant hypomethylation of SLC17A1 in medulloblastoma than that in healthy individual | ||||
Studied Phenotype |
Medulloblastoma [ICD-11:2A00.10] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 2.80E-69; Fold-change: -0.437787629; Z-score: -3.417360823 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Myxopapillary ependymoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
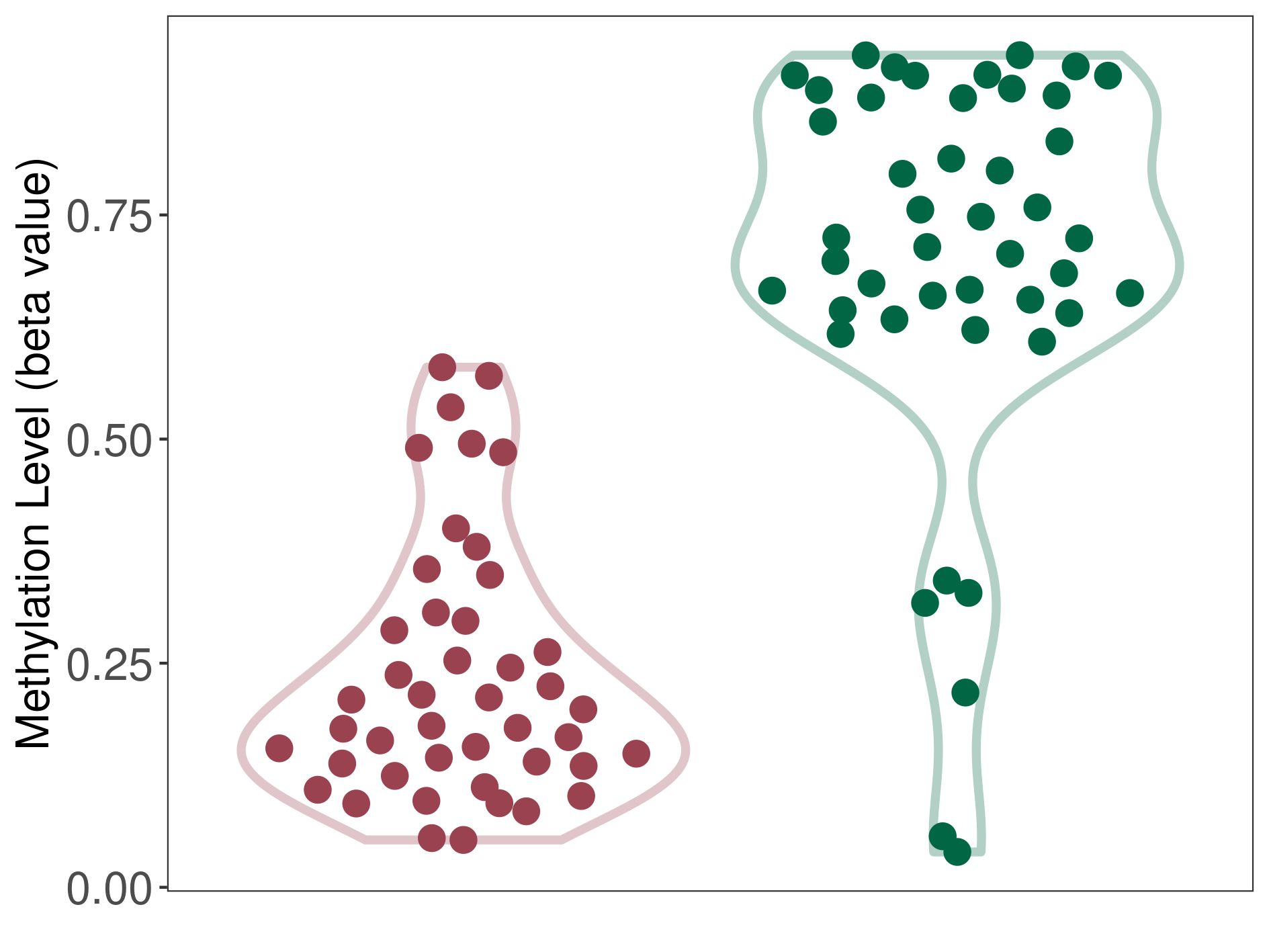
Significant hypomethylation of SLC17A1 in myxopapillary ependymoma than that in healthy individual | ||||
Studied Phenotype |
Myxopapillary ependymoma [ICD-11:2A00.5] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 1.28E-18; Fold-change: -0.534586257; Z-score: -2.414801379 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Obesity |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Significant hypomethylation of SLC17A1 in obesity than that in healthy individual | ||||
Studied Phenotype |
Obesity [ICD-11:5B81] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 4.11E-12; Fold-change: -0.424455702; Z-score: -2.919992715 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||

|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Peripheral neuroectodermal tumour |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Significant hypomethylation of SLC17A1 in peripheral neuroectodermal tumour than that in healthy individual | ||||
Studied Phenotype |
Peripheral neuroectodermal tumour [ICD-11:2B52] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 8.63E-05; Fold-change: -0.359980178; Z-score: -1.510839483 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||

|
 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Uterine carcinosarcoma |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
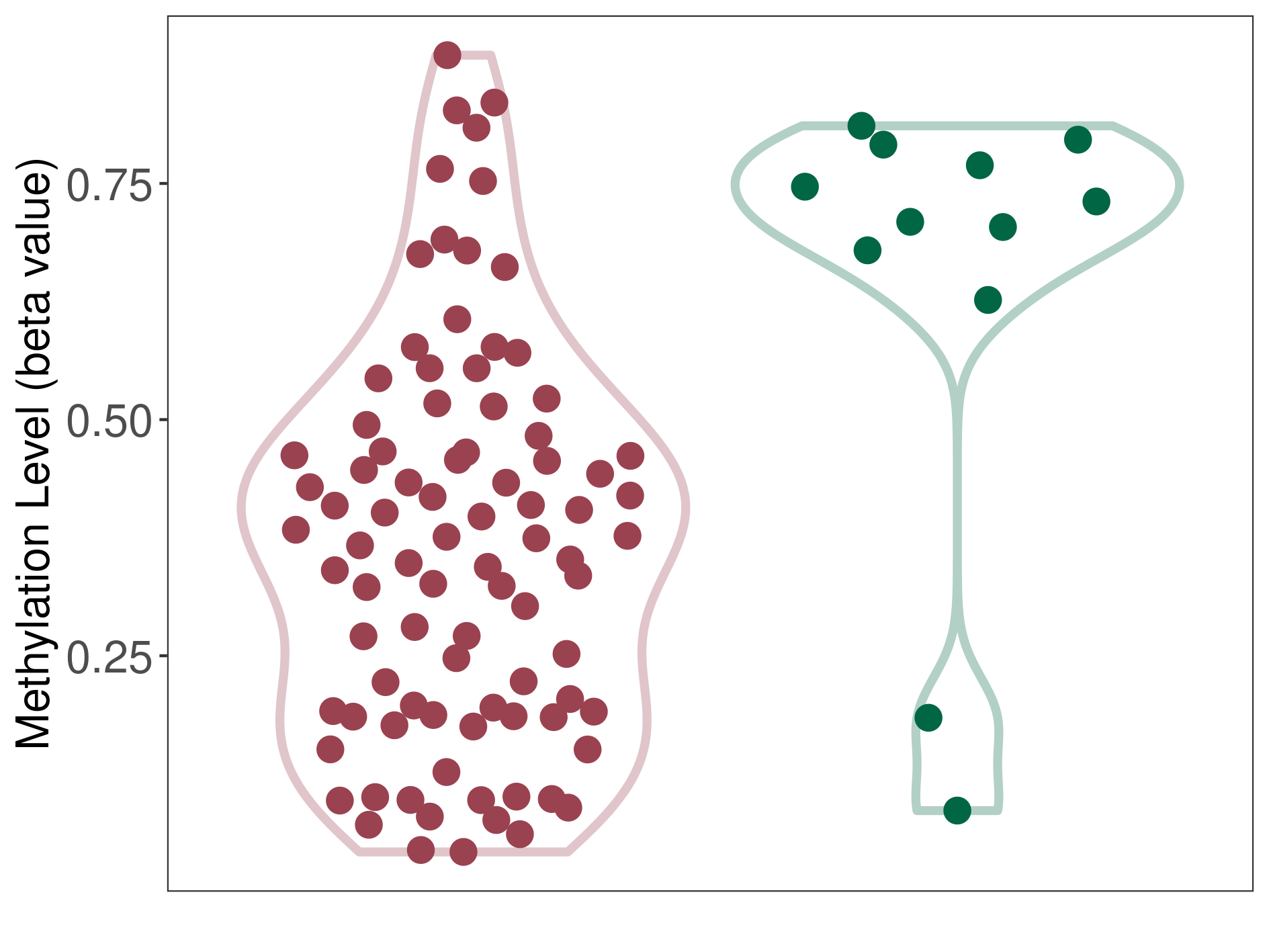
Significant hypomethylation of SLC17A1 in uterine carcinosarcoma than that in healthy individual | ||||
Studied Phenotype |
Uterine carcinosarcoma [ICD-11:2B5F] | ||||
The Methylation Level of Disease Section Compare with the Healthy Individual |
p-value: 0.002467143; Fold-change: -0.349579554; Z-score: -1.451803593 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
|
|||||
|
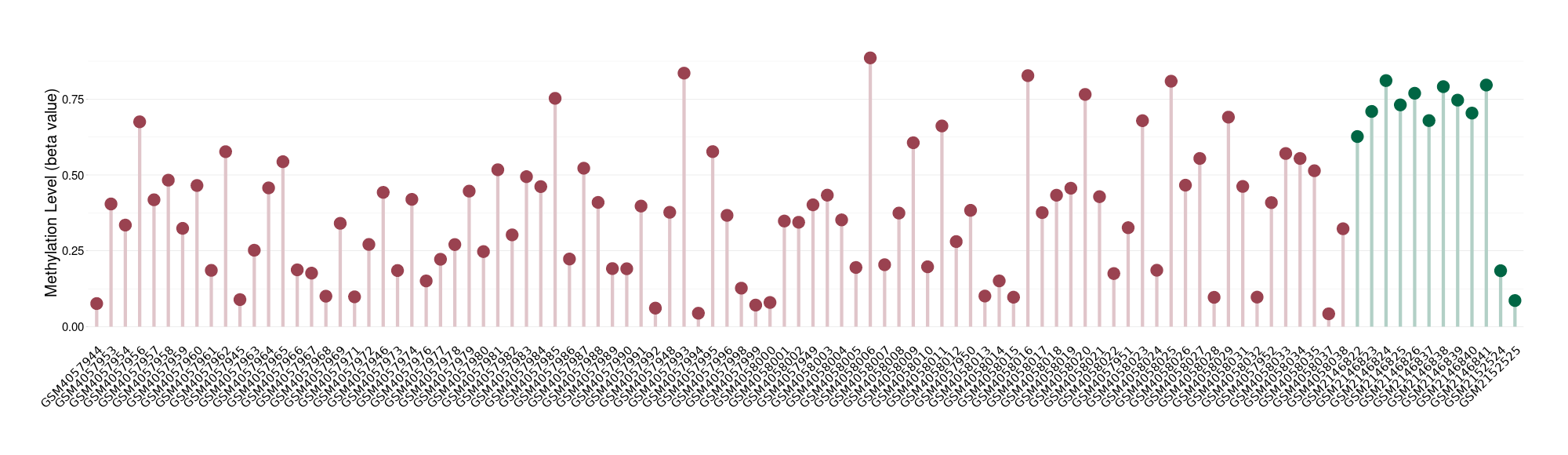 Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
Please Click the above Thumbnail to View/Download
the Methylation Barchart for All Samples
|
||||
|
Lung cancer |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
Significant hypomethylation of SLC17A1 in lung cancer than that in adjacent tissue | ||||
Studied Phenotype |
Lung cancer [ICD-11:2C25] | ||||
The Methylation Level of Disease Section Compare with the Adjacent Tissue |
p-value: 0.000529421; Fold-change: -0.312216056; Z-score: -2.562136049 | ||||
|
DT methylation level in the diseased tissue of patients
DT methylation level in the normal tissue of healthy individuals
DT methylation level in tissue other than the diseased tissue of patients
|
|||||
|
microRNA |
|||||
|
Unclear Phenotype |
1 Epigenetic Phenomena Related to This Phenotype | Click to Show/Hide the Full List | |||
|
Epigenetic Phenomenon 1 |
miR-335 directly targets SLC17A1 | [ 10 ] | |||
|
Epigenetic Type |
microRNA | Experiment Method | Microarray | ||
|
miRNA Stemloop ID |
miR-335 | miRNA Mature ID | miR-335-5p | ||
|
miRNA Sequence |
UCAAGAGCAAUAACGAAAAAUGU | ||||
|
miRNA Target Type |
Direct | ||||
|
Experimental Material |
Multiple cell lines of human | ||||
If you find any error in data or bug in web service, please kindly report it to Dr. Yin and Dr. Li.
